 
Estimation of P-values for global alignments of protein sequences. Selection of a representative set of structures from Brookhaven Protein Data Bank. Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology. Identity and similarity values are often used to assess whether or not two sequences share a common ancestor or function. Thus, it is incorrect to claim that “two sequences are 55% homologous” - we use percent identity or similarity to quote numbers, which we will talk about in the next lesson. Note that homology is a binary qualitative measure - either two sequences are homologous or not. Today, we use the term homology is used to characterize biological species that share a common evolutionary ancestory. All Answers (3) you could directly go to one of the domain databases like pfam and inspect your domains across species, but considering 5 sequences, you can manually retrieve the sequences for. Compare the human amino acid sequence with each of these five animals by counting thenumber of times an amino acid in that animal’s cytochrome c is different from the aminoacid in that. 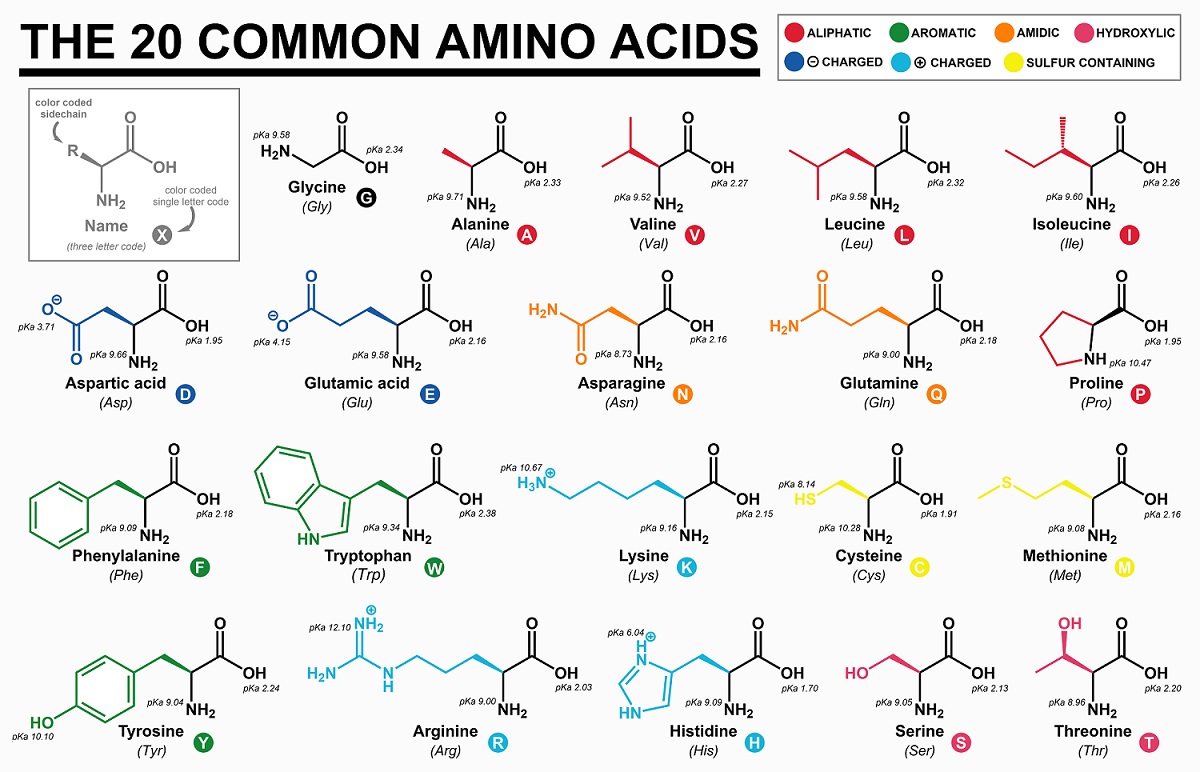
Richard Owen, an English biologist who lived from 1804 to 1892, introduced the term homology, stating that it is “the same organ in different animals under every variety of form and function.” Objects are homologous when they share an evolutionary ancestor. number of equivalenced positions excluding overhangs.It differs, however, from the cytochrome c of rhesus monkeys by 1 amino acid, from that of horses by 11 additional amino. For example divide the number of identities by: chimpanzees, the protein molecule called cytochrome c, which serves a vital function in respiration within cells, consists of the same 104 amino acids in exactly the same order. Having got the alignment by some method, there are many different ways of calculating percentage identity (PID). Sequence identity is the amount of characters which match exactly between two different sequences 
Identity and similarity are very often used interchangeably (for nucleotide sequence or in the context of graph theory) Sequence Identity 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
